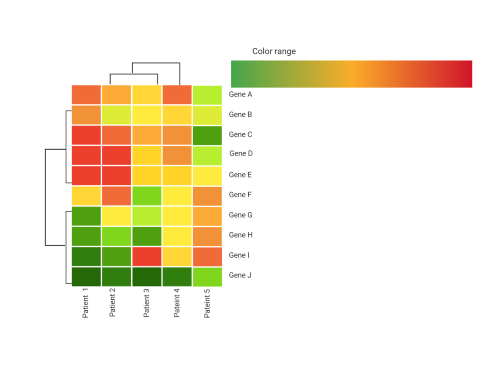
Can we integrate genomics and other molecular features to improve the prognosis of CMML?
There exist at least 10 prognostic models that have been reported to predict survival in CMML making it the cancer with the largest number of prognostic models per-capita. However, despite the addition of several novel models that incorporate genomics to predict survival in CMML, no consensus model exists. Further, we have demonstrated that all models have comparatively modest predictive power in a large international database of over 1800 patients. Therefore, there exists a critical need to credential existing models in a uniform molecularly and clinically annotated database to identify a consensus approach. To address this, we have established a collaboration, through the Myelodysplastic Syndrome International Working group, that will uniformly sequence and clinically annotate 1000 diagnostic CMML samples from 22 centers worldwide. This data set will serve to bench mark existing prognostic models and, if necessary, development of a novel consensus model.
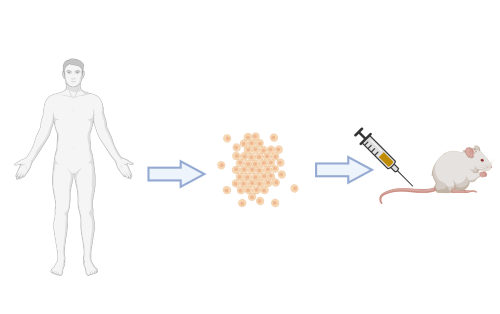
Can we effectively model CMML preclinically?
One central limitation to the development of effective therapeutic approaches for CMML has been the absence of preclinical CMML model systems. This includes the lack of cell lines or robust liquid cell culture models of primary CMML. A number of groups have attempted to develop genetically engineered mouse models (GEMMs) of CMML based on modeling frequently occurring somatic mutations. While these models have been very helpful for mechanistic studies aimed at understanding how these mutations alter hematopoiesis, none of these models accurately recapitulate the phenotypic or genetic heterogeneity of CMML. To this end, our group, in collaboration with the Abdel-Wahab lab, have generated the first robust patient derived xenograft (PDX) models of CMML that accurately recapitulate the genetic and phenotypic characteristics of the human condition. Our current focus is to continue to characterize the fidelity of this model and explore its application towards the identification of novel therapeutics.
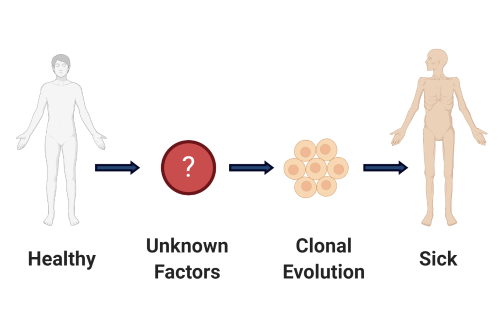
Can we develop strategies to prevent the development or progression of CMML in patients at risk?
In current clinical CMML practice, patients are treated once disease progression or disease-related symptoms are identified. However, several lines of clinical evidence from our group and others suggests that early intervention may be beneficial in some CMML subsets. Therefore, identifying the factors that lead to clonal evolution and disease progression is important to develop therapeutics to prevent this dismal natural history. Given our observation demonstrating that cytokines and other inflammatory mediators are relevant to disease progression in CMML, we aim to provide evidence to support an alternative treatment strategy whereby clinically asymptomatic patients would be treated with relevant inhibitors resulting in stabilization of disease.
Can we identify novel therapeutic vulnerabilities in CMML?
Hypomethylating agents are FDA approved for use in CMML. However, this approval was based on the assumption that CMML was a subtype of MDS which is the disease in which hypomethylating agents were rigorously clinically studied. It is now known that CMML is a distinct entity and, as such, no evidence is available to suggest that these therapies augment the natural history of CMML. Further, clinical trials for CMML are rare providing limiting opportunities for patients with this lethal cancer. Our laboratory aims to identify novel, clinically tractable, therapeutic targets for CMML. Examples of clinical trials which arose, in part, from preclinical work from our laboratory include ruxolitinib for CMML and lenzilumab for CMML.